(ten Tusscher KH, Noble D, Noble PJ, Panfilov AV. A model for human ventricular tissue. Am J Physiol Heart Circ Physiol. 2004;286(4):H1573-89)
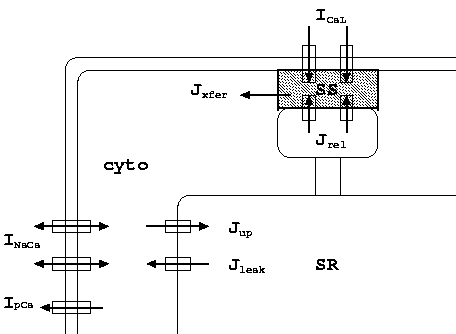
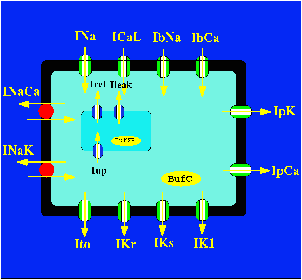
The model is based on human based experimental data for the individual ionic currents (INa, Ito, IKr, IKs, IK1, ICaL), action potential shape, duration, restitution, and ionic concentration changes.
A Java applet of the human ventricular cell model. The applet was developed by Flavio Fenton and Elizabeth Cherry from Hofstra University .